Virtual Brain Project
This project is developing a comprehensive framework for modelling the brain using computational methods built on bioengineering principles.

Our aim
The human brain is the most complex and the least well-understood organ of the human body. Bioengineering contributions to brain research have been relatively few so far and we would like to address this by developing a comprehensive framework for modelling the brain using computational methods built on bioengineering principles. The projects outlined below include studies of the cerebral circulation and nutrient transport, neurovascular coupling, biophysical aspects of brain injury, and mapping studies that link neural models of motor control to the visceral organs, both for the autonomic and somatic motor systems.
Research areas
Advanced AI-driven image processing
Due to the recent breakthroughs in artificial intelligence technologies, techniques for subject-specific digitalisation enable us to reconstruct the human brain in-silico from MRI sequences.We explore the anatomical variations of the human brain by deep learning and dimensionality reduction techniques, towards a major understanding of its constitutive structures and its diversity among the population. Such insights aim to improve the statistical models used to investigate pathological scenarios as mild cognitive impairment (MCI) and Alzheimer’s Disease (AD).
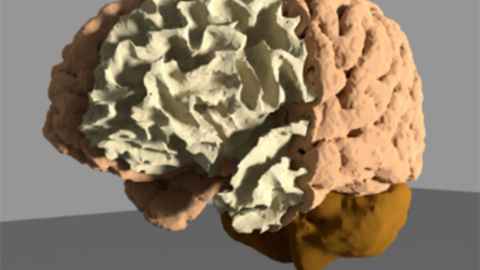
By means of this in-silico representation of the human brain, computational neurovascular simulations are available for research and future clinical usage. This fusion of AI image processing with computational models in a unified environment, yields novel tools to (i) study the fundamental processes in the neurovascular coupling; (ii) development of novel biomarkers for early diagnosis of MCI and AD; and (iii) study the association between cardiovascular patterns and pathological brain conditions.
Research Lead: Gonzalo Maso Talou
Developing an in-silico platform for modelling cardiovascular system
Funded under the ‘Science for Technological Innovation’ (SfTI), we built a novel in-silico CellML model of the entire cardiovascular system using a bond graph formulation that runs faster than real-time on a desktop computer and is designed for use in clinical settings when the speed of response is important. This computational framework is able to simulate the blood flow through the vascular system including the cerebral arteries. The bond graph formulation makes it straightforward to extend the model to include various tissue exchange mechanisms and to incorporate tissue and cellular parameters that characterise various chronic diseases.
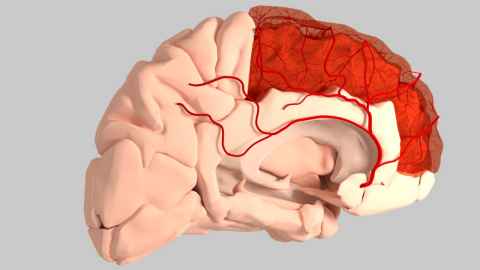
This model can also include self-regulating and metabolic dynamics in a simple way and at a low computational cost. Cerebral autoregulation is a precise system involving vasodilation and vasoconstriction in a network of collateral vessels. By adding metabolic models to the bond graph model we would be able to simulate cerebral autoregulation, which is a feedback mechanism driving an appropriate blood supply into the cerebral vasculature depending on the oxygen demand by the brain. There are also many other physiological mechanisms that can be added to this system such as baroreflex regulation, respiratory control system, autonomic nervous system, etc.
Research Lead: Soroush Safaei
Multi-scale modelling of neurovascular coupling
The brain is critically dependent on a continuous supply of blood to function. Therefore, the cerebral vasculature is endowed with neurovascular control mechanisms that assure that the blood supply of the brain is commensurate to the energy needs of its cellular constituents. The regulation of cerebral blood flow (CBF) during brain activity involves the coordinated interaction of neurons, glia, and vascular cells. Thus, whereas neurons and glia generate the signals initiating the vasodilation, endothelial cells, pericytes, and smooth muscle cells act in concert to transduce these signals into carefully orchestrated vascular changes that lead to CBF increases focused to the activated area and temporally linked to the period of activation.
Neurovascular coupling (NVC) is disrupted in pathological conditions, such as hypertension, Alzheimer’s disease, and ischemic stroke. Consequently, CBF is no longer matched to the metabolic requirements of the tissue. This cerebrovascular dysregulation is mediated in large part by the deleterious action of reactive oxygen species on cerebral blood vessels. Mathematical modelling can be a useful tool for investigating the contribution of various signalling pathways towards NVC and for analysing the underlying cellular mechanisms. It can also provide insight into the interaction of the cellular-level and macro-scale phenomena studied in-silico and a deeper understanding of experimental results.
Given that neuronal activity is closely linked to cerebral blood vessels, we propose a neurovascular approach to bridge the gap between vascular and neurodegenerative disorders and tackle Alzheimer’s Disease. By employing computational modelling, here we investigate the vascular regulatory mechanisms for NVC, an essential process in the delivery of oxygen and nutrients to the brain, as well as in the clearance of by-products of brain metabolism. The ultimate goal of this project is to facilitate the development of new therapeutic strategies for brain disorders in which the above process fails.
Research Lead: Soroush Safaei
Computational neuroscience
Understand the underlying mechanisms of brain physiology and neural activity from subcellular to whole-CNS, and study the nerve-organ interaction using experimental, computational, and mathematical means.
- Computational models of interoception, body regulation, and forecasting. Specifically, the interoceptive inference and how bodily states are regulated by autonomic reflexes that are forced by predictions from deep internal (generative) models of internal and external milieu. This falls in the idea of the “Bayesian Brain” in the context of interoception which is really perception and integration of autonomic, hormonal, visceral, and immunological signals (i.e., sense of the body from within). This also tightly relates to the role of interoception in the sense of embodied self and feelings (importance of functional anatomy of the emotional brain) - this can range from psychophysiological conditions such as autism, anxiety, and depression through to consciousness.
- Synaptic plasticity and consolidation and their role in receptive field development. For example, what is the mechanism behind orientation selectivity during learning in the visual cortex? Additionally, the relation between the type of coding and the type of connectivity in different brain areas. This will include: developing biologically-driven models (currently working with active inference, variational free energy, and predictive coding, etc.), reproducing experimental findings (e.g., images, videos, slice experiments, ephys, etc.) with models.
- Extending the idea of synaptic plasticity & interoceptive inference, how does the brain create the “cognitive map” and the generative model? Where is this model? How does space and time influence the model and, consequently, our perception of the world? How does this model (or its parameters) adapt to [future] perturbations (internal and external) and thus maintain homeostasis (or allostasis)?
Research Lead: Gonzalo Maso Talou
Multi-scale modelling of traumatic brain injury
Traumatic brain injury (TBI) is a growing health and socio-economic problem and yet its mechanism is still poorly understood. Proper management of TBI is essential to avoid short-term and long-term risks ranging from brain haemorrhage to dementia. Therefore, accurate quantification of how mechanical energy from the external impact is transferred into the brain and its cells is critical for understanding the mechanisms of TBI and developing effective diagnostic and treatment options. Our multidisciplinary project aims to achieve this. Multiscale computational models and cell mechanical devices are being combined with cutting edge neuroscience to identify injury thresholds for TBI and possible therapeutic targets for effective treatment. This will provide the basis for developing a comprehensive system made up of wearable sensors and handheld devices that can be used in measuring head impact and providing real-time diagnosis and treatment options.
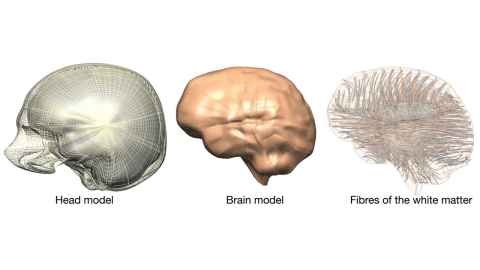
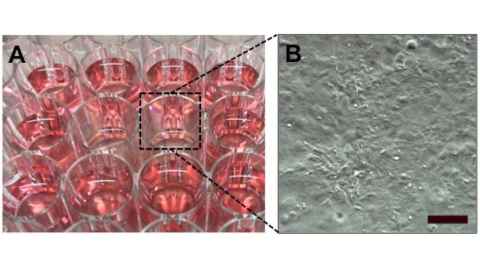
This project is an interdisciplinary collaboration between two research centres at the Universityof Auckland, (The Auckland Bioengineering Institute and the Centre for Brain Research) as well as with a brain concussion sensor company CSx. We have already developed an accurate model of the human brain (see Figure 1) using the Auckland Bioengineering Institute’s own open source software and devices that can apply impact load to cells (see Figure 2). The Centre for Brain Research at the Faculty of Medical and Health Sciences have been at the forefront of neuroscience research and has developed novel 2D and 3D human brain cell culture systems (see Figure 3). This state-of-the-art neuroscience will becombined with:
- Wearable sensors (from CSx) measuring actual brain concussionfrom NRL rugby players
- High fidelity computational models of the brain ofthose rugby players
- Advanced cell mechanical devices to apply in-vivo likesignals to brain cells directly
Once combined, we will be able to recreate the cellular micro environment of brain cells during concussion to identify key mechanisms that cause damages to the brain. This will be used to develop potential therapies for TBI.
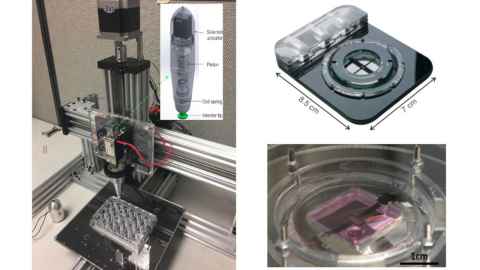

Research Lead: Vickie Shim
Using MRI to assess blood-brain barrier permeability
The blood-brain barrier (BBB) regulates the exchange of nutrients, signalling molecules and cells between the vascular system and brain tissue. Disruption of the BBB has been associated with neurological conditions such as stroke, brain cancers and neurodegenerative disorders.
Recently, subtle damage to the BBB measured in-vivo with MRI has been found in mild cognitive impairment (MCI) and Alzheimer’s Disease (AD). This raises the possibility that BBB permeability can serve as an early biomarker of disease that could indicate now, who will progress from MCI to AD. Such biomarkers will enable us to develop interventions and treatments that slow down or prevent the development of dementia.
We are developing non-contrast based MRI protocols and data analysis methods to develop a non-invasive MRI marker of BBB permeability that can be used for early disease detection. By understanding BBB dysfunction in MCI and AD, this method will allow in-vivo BBB research to be carried out in a wide range of applications such as normal ageing, stroke, Parkinson’s disease and motor neurone disease.
Research Lead: Vinod Suresh
Members
Primary contact
Academics
Finbar Argus
Soroush Safaei
Vickie Shim
Vinod Suresh
Gonzalo Maso Talou
Professionals
Chinchien Lin
Shan Su
Students
Cameron Apeldoorn
Stephen Creamer
Sergio Dempsey
Harshil Magan
Robyn May
Alireza Sharifzadeh
Jiantao Shen
Gurleen Singh
Collaborators
Pablo Blanco (HeMoLab)
Oscar Camara (Universitat Pompeu Fabra)
Gunnar Cedersund (Linköping University)
Maurice Curtis (CBR)
Michael Dragunow (CBR)
David Dubowitz (CAMRI)
Samantha Holdsworth (CBR)
Victoria Low (CBR)
Tracy Melzer (University of Otago)
Johanna Montgomery (FMHS)
Catherine Morgan (CBR)
Lucas Muller (University of Trento)
Thomas Park (CBR)
Julian Paton (FMHS)
Reece Roberts (University of Auckland)
Samuel Rosset (University of Auckland)
Mark Sagar (University of Auckland)
Miriam Scadeng (CBR)
Alan Wang (University of Auckland)
Funding partners
- Neurological Foundation of New Zealand
- New Zealand Ministry of Business Innovation and Employment (MBIE)
- NIH Common Fund
- The Aotearoa Foundation
- The Li Ka Shing Foundation
- The Marsden Fund
International links
- Brazil: HeMoLab
- Italy: University of Trento
- Norway: Norwegian University of Science and Technology
- South Korea: Seoul National University
- Spain: Universitat Pompeu Fabra
- Sweden: Linköping University
- United Kingdom: University of Oxford
- United States: UC Berkeley, University of Michigan